Earlier this year, Columbia University Press published a book entitled “Junk DNA: A Journey Through the Dark Matter of the Genome” by Nessa Carey. A popular science book written for a general audience, “Junk DNA” is an exploration of all the ways that DNA can function without coding for proteins. As the author rightly decries, the protein-coding regions of our DNA have gotten almost all of the attention for far too long.
Of course, the snubbing of noncoding DNA was totally understandable. Genes and their products were the first aspects of DNA that were discovered and the easiest to understand. In fact, many of the functions of noncoding DNA are to modulate the expression of genes, so without an understanding of how genes work, there would be no way to discover what these regulatory regions do. Furthermore, the field of molecular genetics got its start by studying organisms with much smaller and simpler genomes, things like bacteria and viruses. These organisms have almost no “junk DNA,” so its importance was largely overlooked for decades.
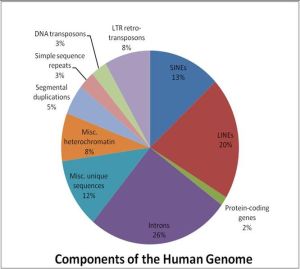
Only around 1.5% of the human genome codes for proteins and this book is all about the rest of it – the 98.5% of the genome that we only now are starting to understand. As such, this book is an excellent choice for science students and others interested in learning more about what our chromosomes really contain besides a few genes here and there.
An important caveat that I feel compelled to note for those planning to read it, this book employs the term “junk DNA” very broadly, applying it to any stretch of DNA sequence that does not code for protein. Even genes that encode RNA molecules are referred to as junk in this book. While this may be common practice among non-specialists, in my 17 years as a molecular biologist, I’ve not heard a scientist refer to promoters, introns, terminators, centromeres, or telomeres as “junk.” The functions of those elements have been known for quite a long time. Indeed, even back in the 1970s, promoters and introns were appreciated as essential components of genes. An expert molecular biologist herself, Dr. Carey surely knows this and the broad use of the “junk DNA” label appears to be a dramatic authorial choice and, as such, it works.
Here is a partial list of the kinds of “dark matter” within the human genome that are explored in “Junk DNA”
- Introns – the regions within a gene that do not encode for part of the protein and are instead “spliced out” of the mRNA before it is translated to protein.
- ribosomal RNA – The workhorses of protein synthesis are made of more RNA than protein and the genes that encode these rRNAs have been repeatedly duplicated and are present throughout our genome in high numbers.
- Centromeres – the structural DNA sequence marking the “center” of the chromosomes for proper segregation during mitosis
- microRNAs – the most recently discovered kind of RNA, which has revolutionized our understanding of gene expression control. Thousands of genes for crucial miRNAs have been found.
- Viral genomes – some viruses, especially retroviruses, implant their genetic material into our own and sometimes it remains there and is passed on for millions years.
- Telomeres – the extreme ends of the chromosomes with a special structural sequencing for maintaining or capping.
- promoters, enhancers, silencers – regulatory sequences to which factors bind to influence expression of a gene
- Highly repetitive DNA – the true “dark matter” DNA whose nature and function is the most mysterious.
- small nuclear RNAs – these RNAs regulate other genes often by affecting how their mRNA is spliced into a final product. They can even do this differently in different cells, allowing us to make slightly different versions of a protein in different tissues.
There is also considerable coverage of exciting developments in the field of epigenetics, the subject of Dr. Carey’s previous book.
“Junk DNA: A Journey Through the Dark Matter of the Genome” is an engaging and fast-paced read. Each chapter is dedicated to a specific type of noncoding DNA and its functions are discussed, often in great detail. In many cases, the author takes us through some of the key experiments that helped shape our understanding, so that the readers get a sense of how we know what we know. In addition, Dr. Carey explains the genesis of various diseases that can result from dysfunction in noncoding DNA, including an appendix dedicated sole to that. This helps underscore the importance of these long-mysterious parts of our genome.
This book is very well-written and fun to read. One of my favorite aspects of Dr. Carey’s writing style is her frequent use of clever analogies. For example, she compares the expression of some noncoding RNAs to the stray wood chips and shavings that result from chopping down a tree. Those “stray RNAs,” like the wood shavings, aren’t made deliberately and they don’t serve any function, but, at the same time, they don’t really cause any problems or cost anything either. I will definitely be using that analogy when I teach this material to my biology major students.
High school and college students will greatly benefit from reading this book and gain important perspective on the richness of the human genome and the mysterious aspects of it that await thorough understanding. Any reader with an interest in biology or medicine will enjoy this book as well and it is widely accessible. You need not be an expert to understand the content and absolutely everyone that reads it, even specialists, will learn things.
While I unequivocally endorse and recommend this book, there is one point on which I may hold a different view than the author. There is a sense in this book that, since we have discovered functions for much of the previously called “junk,” that we will eventually find functions for all of it. I don’t think that is the case.
First of all, it has been known for some time that DNA elements can do nothing more than support their own proliferation. Some do this through expression, but not all do. Transposons and retrotransposons behave almost like organisms, copying themselves and moving through our genome. For all intents and purposes, these really are junk, and the book almost, but not quite, admits this. There isn’t an easy mechanism for us to get rid of this junk and so it has accumulated over time. That these highly repetitive elements haven’t been eliminated isn’t necessarily evidence for their having a function; it can merely be evidence of the success of their self-propagating nature. While it is true that they have contributed positively to genome evolution, as a whole, and that individual elements can develop functions serendipitously, large stretches of this repetitive DNA does nothing (or does nothing good), even if it is sometimes accidentally transcribed.
Secondly, this book, as other writing about noncoding DNA, is filled with examples of how junk DNA can be involved in the genesis of diseases. It appears that the underlying assumption is that, since these elements can cause disease when they go awry, they must have important functions in the normal/healthy state. This is not always true, including in the examples discussed in this book. Sometimes the normal/healthy state is that the genetic element does nothing at all. Then, it causes a disease state by making too many copies of itself or jumping into, or too close to, a gene.
In my view, the author says too many times, “we don’t yet know what the proper function of this DNA sequence is.” Ultimately, It may not have one.
Nevertheless, I very much enjoyed reading this book and I think it is an excellent choice for any nonspecialist who would like a captivating, thorough, and up-to-date introduction to the dark recesses of our genome. Molecular biology has long had a gene-centric view of our DNA and this book does an excellent job of getting the casual reader up to speed about the bigger picture of our wonderful and messy genome.
[Update: My new book, “Human Errors” devotes a whole chapter to our sloppy genome! Check it out!]
-NHL








Leave a comment